download
Optimizing PCR Amplification in Single-cell RNA-seq
Learn how real-time PCR control streamlines single-cell RNA-seq library prep while preserving data quality across variable inputs by automatically stopping amplification at the optimal cycle.
Traditional thermocyclers require you to choose a cycle number before you know exactly how your samples will behave.
Choosing a fixed cycle number poses a challenge for PCR amplification in single-cell RNA-seq library preparation. It means estimating amplification based on the number of cells and the RNA content per sample.
The problem is that different inputs require different cycling conditions. So you split samples across multiple runs, quantify libraries between PCR steps, and hope you haven’t over- or under-amplified.
The result? Extra hands-on time, workflow complexity, reagent waste, and libraries that reflect amplification artifacts rather than biology.
In this application note, n6 demonstrates how icon96™ with AutoNorm™ improves library preparation by eliminating the need for manual normalization. By replacing fixed-cycle amplification with real-time PCR control, each reaction is monitored individually and automatically stopped at the optimal point in the amplification curve, removing the need for pre-quantification or batching by cell number and consistently delivering sequencing-ready libraries.
Discover how, compared to a standard 10x Genomics workflow, n6’s icon96™ with AutoNorm™ technology maintains single-cell gene expression data quality whilst simplifying your library prep workflow by:
- Simultaneously running samples of input ranges from 500 to 10,000 cells, decreasing the number of thermocycler runs
- Reducing the total number of PCR cycles per sample, minimizing overamplification
- Eliminating pre-index PCR quantification for faster turnaround times
- Reducing hands-on time by 40 to 60%
Download the application note to learn how real-time PCR control can streamline your single-cell RNA-seq workflow and improve data quality.
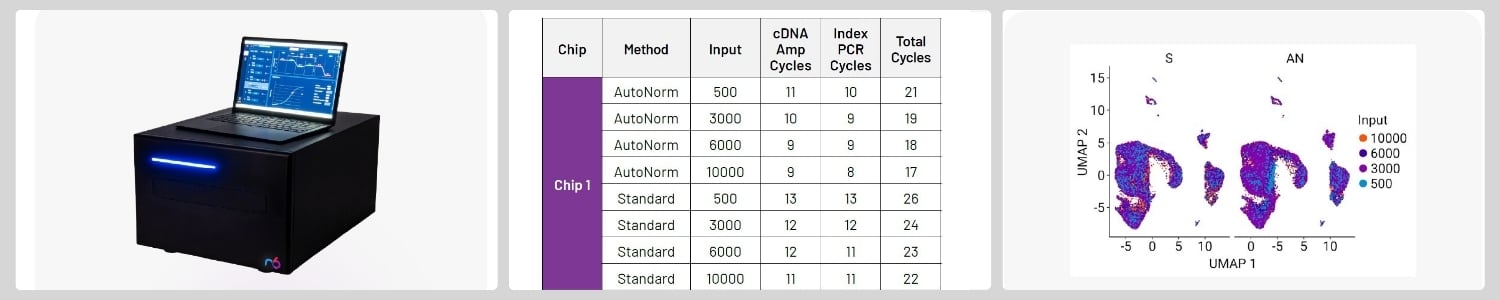
From left to right: n6’s icon96™ real-time thermocycler; table showing PCR cycling comparisons between icon96™ with AutoNorm™ technology and the standard PCR protocol; clustering of samples demonstrating no difference in data quality between the protocols.