download
Reduce Pipetting Variability in Your Lab
Practical strategies to improve accuracy, standardize technique, and strengthen experimental reproducibility
If you want to reduce pipetting variability, you need more than just yearly calibration checks.
Pipetting is fundamental to most life science workflows. When setting up qPCR assays, performing ELISAs, running Western blots, conducting cell-based assays, and performing myriad other analyses, even small-volume inconsistencies can propagate through your experiment, amplifying errors and compromising reproducibility.
Because pipetting feels routine, it’s not often considered a major source of variation. Yet, instrument performance, tip fit, technique, maintenance schedules, and protocol design can all affect your results.
This white paper takes a practical, bench-focused approach to help you reduce pipetting variability across users and experiments.
Inside, you’ll learn how to:
• Choose pipettes based on your workflow and error specifications
• Select tips that seal properly, minimize retention, and prevent batch effects
• Standardize pipetting technique to minimize user-to-user differences
• Implement routine performance verification and smarter calibration intervals
By applying the principles described in this guide, you can improve consistency in your liquid-handling workflows, build confidence in your quantitative results, and make your experiments easier to reproduce.
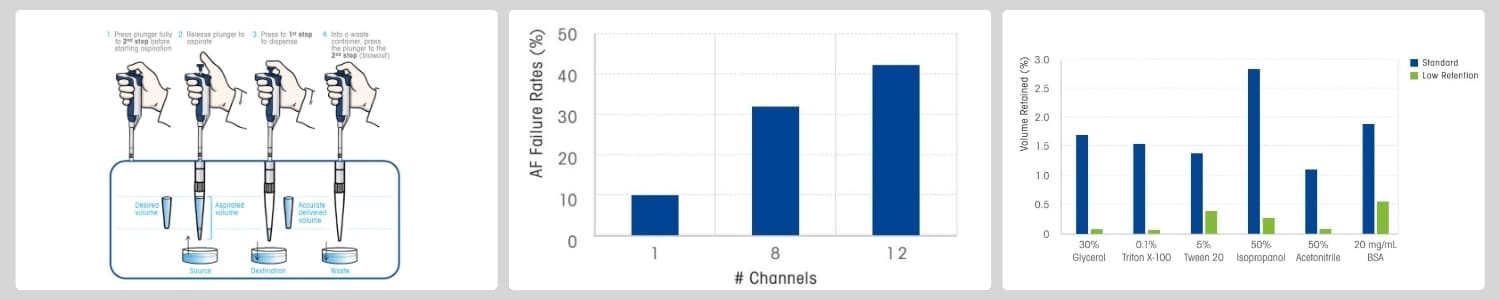
Figure 1. Visual guide to reverse-pipetting technique, failure rate of pipettes is positively correlated with the number of channels, and comparison of low retention tips with standard tips for pipetting different liquids.

